Welcome to GREAP a comprehensive human genomic region sets enrichment analysis platform.
- Introduction
-
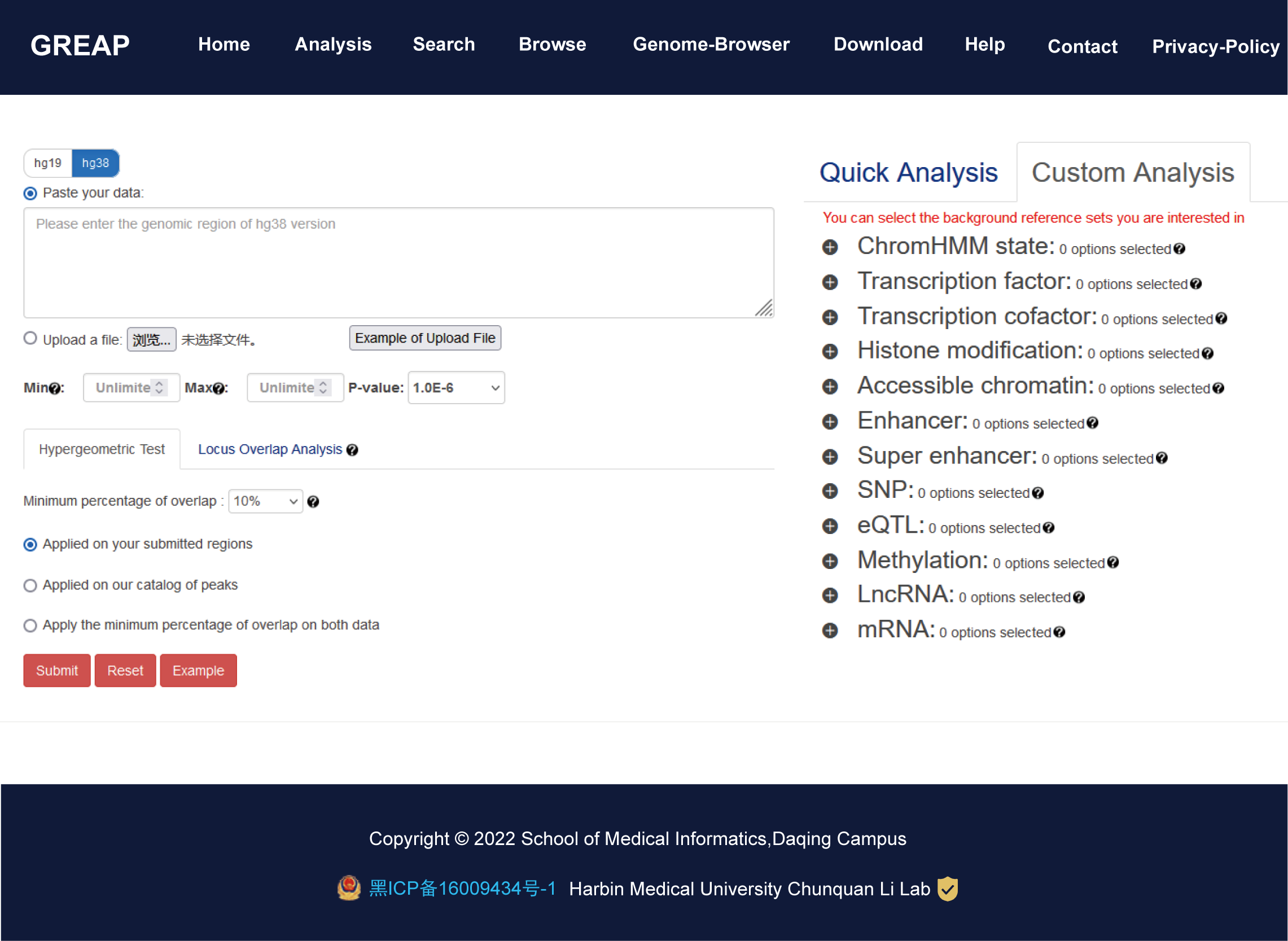
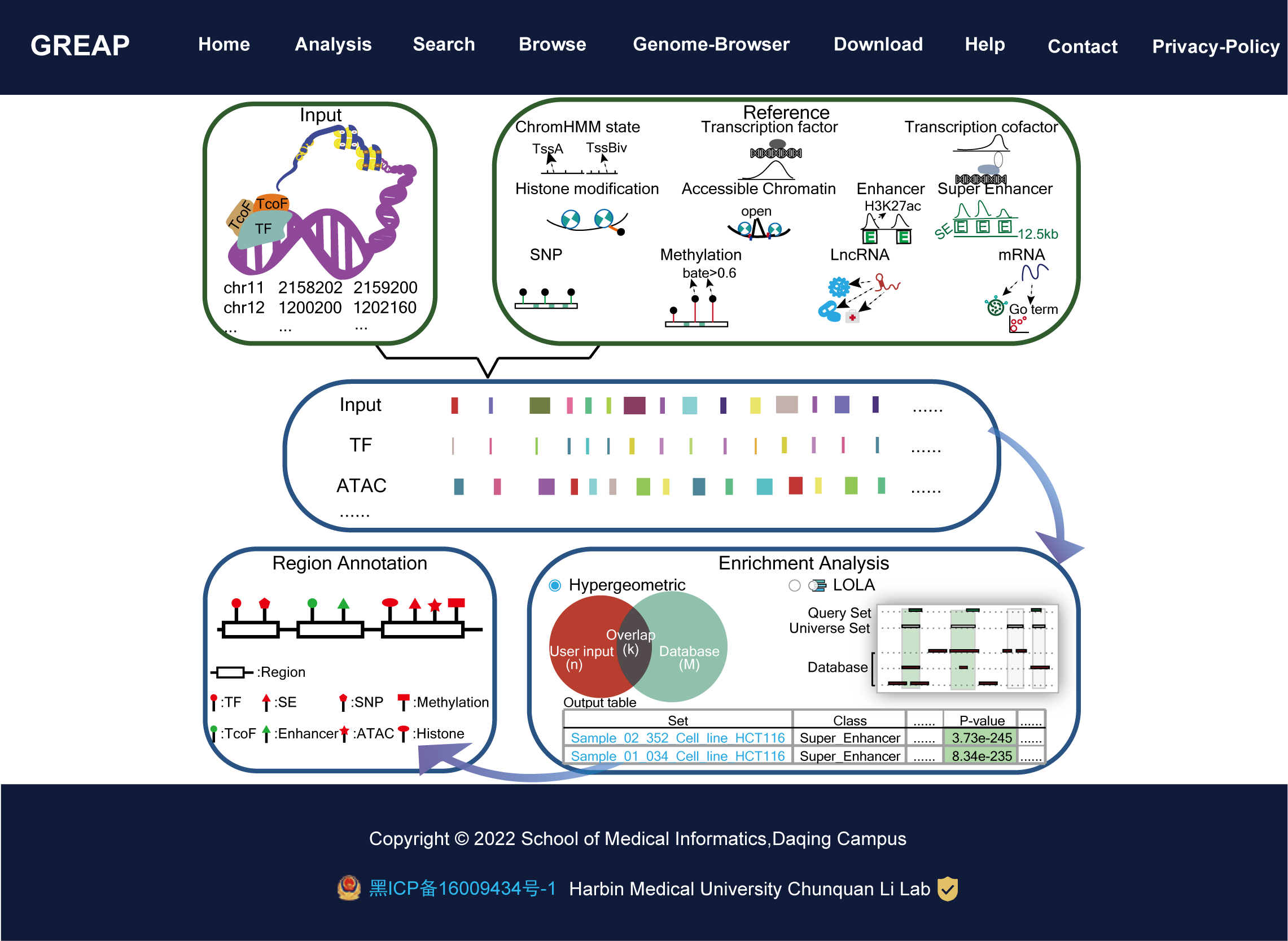
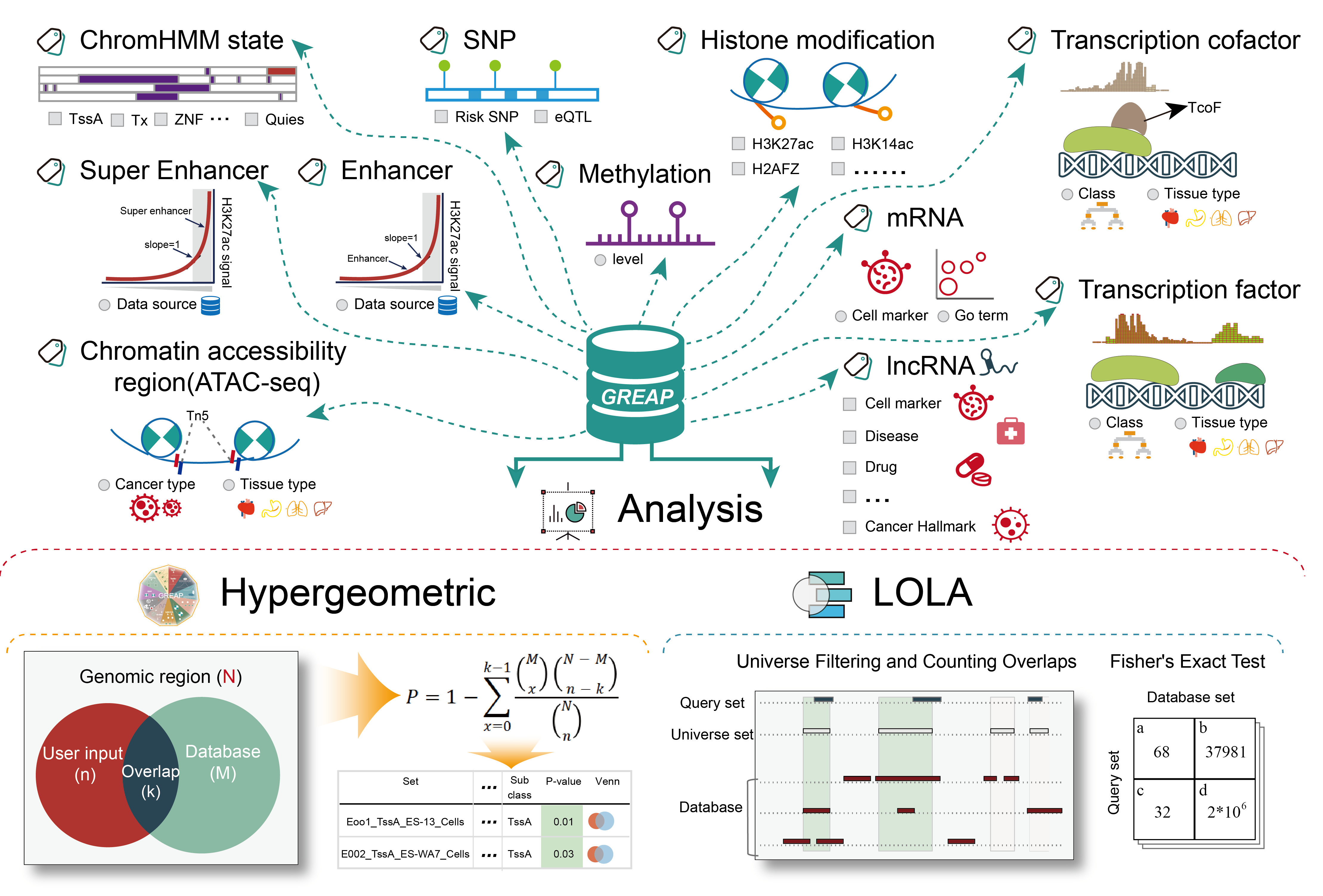
 GREAP aims to document a large number of available resources of human genomic region sets and to provide annotation and enrichment analysis of region lists submitted by users.GREAP supports 85,370 genomic region sets involved in 11 categories (ChromHMM state, TF, TcoF, Histone modification, Accessible Chromatin, Enhancer, Super Enhancer, SNP, Methylation, LncRNA, mRNA) and 454 sub-categories, which covered over 634,681,107 regions. Moreover, we provide two region enrichment analysis methods: (i) Hypergeometric test, a common enrichment analysis method which is frequently cited, (ii) Locus Overlap Analysis (LOLA) uses Fisher's exact test with false discovery rate correction to assess the significance of overlap in each pairwise comparison.
GREAP aims to document a large number of available resources of human genomic region sets and to provide annotation and enrichment analysis of region lists submitted by users.GREAP supports 85,370 genomic region sets involved in 11 categories (ChromHMM state, TF, TcoF, Histone modification, Accessible Chromatin, Enhancer, Super Enhancer, SNP, Methylation, LncRNA, mRNA) and 454 sub-categories, which covered over 634,681,107 regions. Moreover, we provide two region enrichment analysis methods: (i) Hypergeometric test, a common enrichment analysis method which is frequently cited, (ii) Locus Overlap Analysis (LOLA) uses Fisher's exact test with false discovery rate correction to assess the significance of overlap in each pairwise comparison.
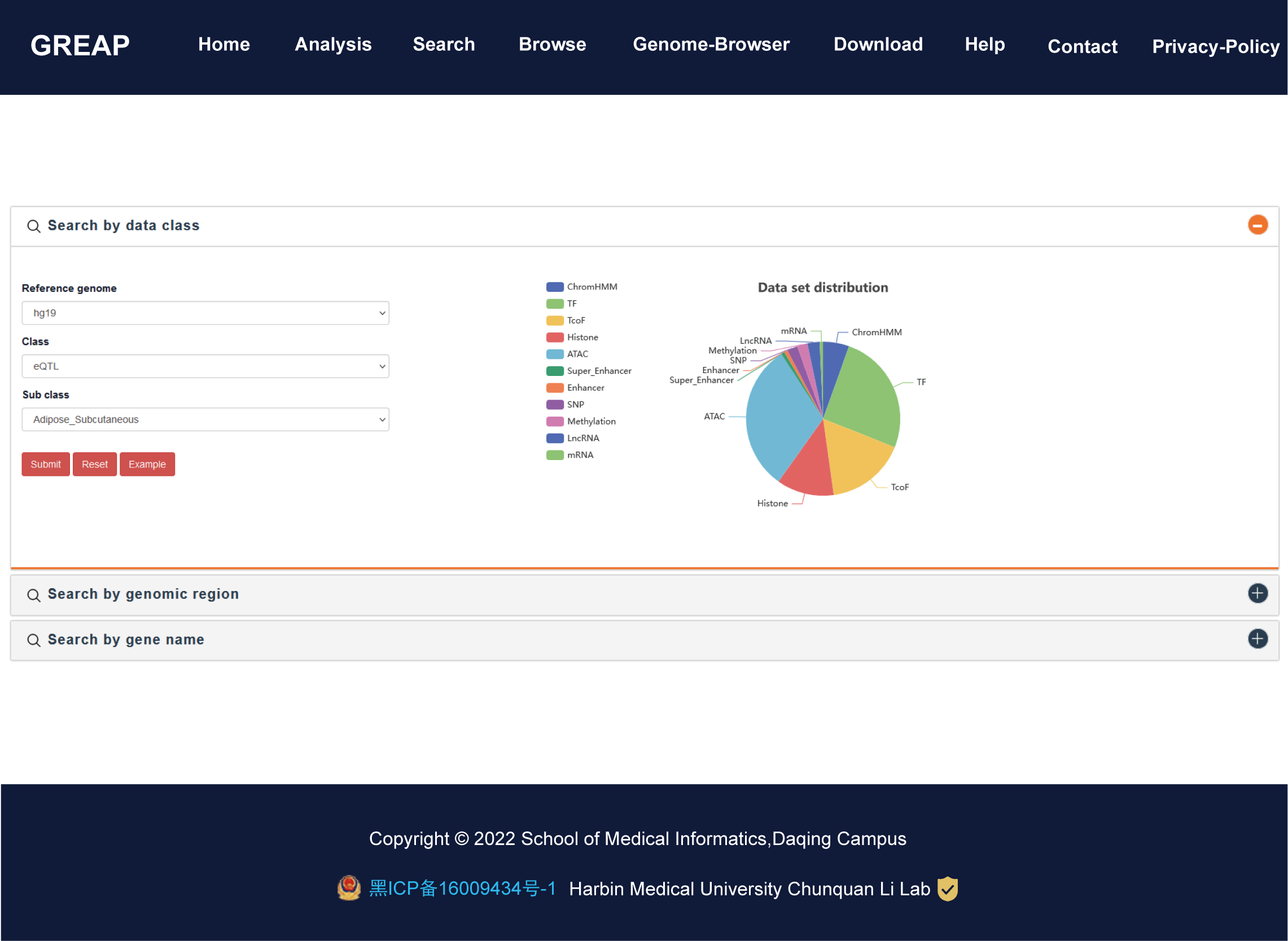
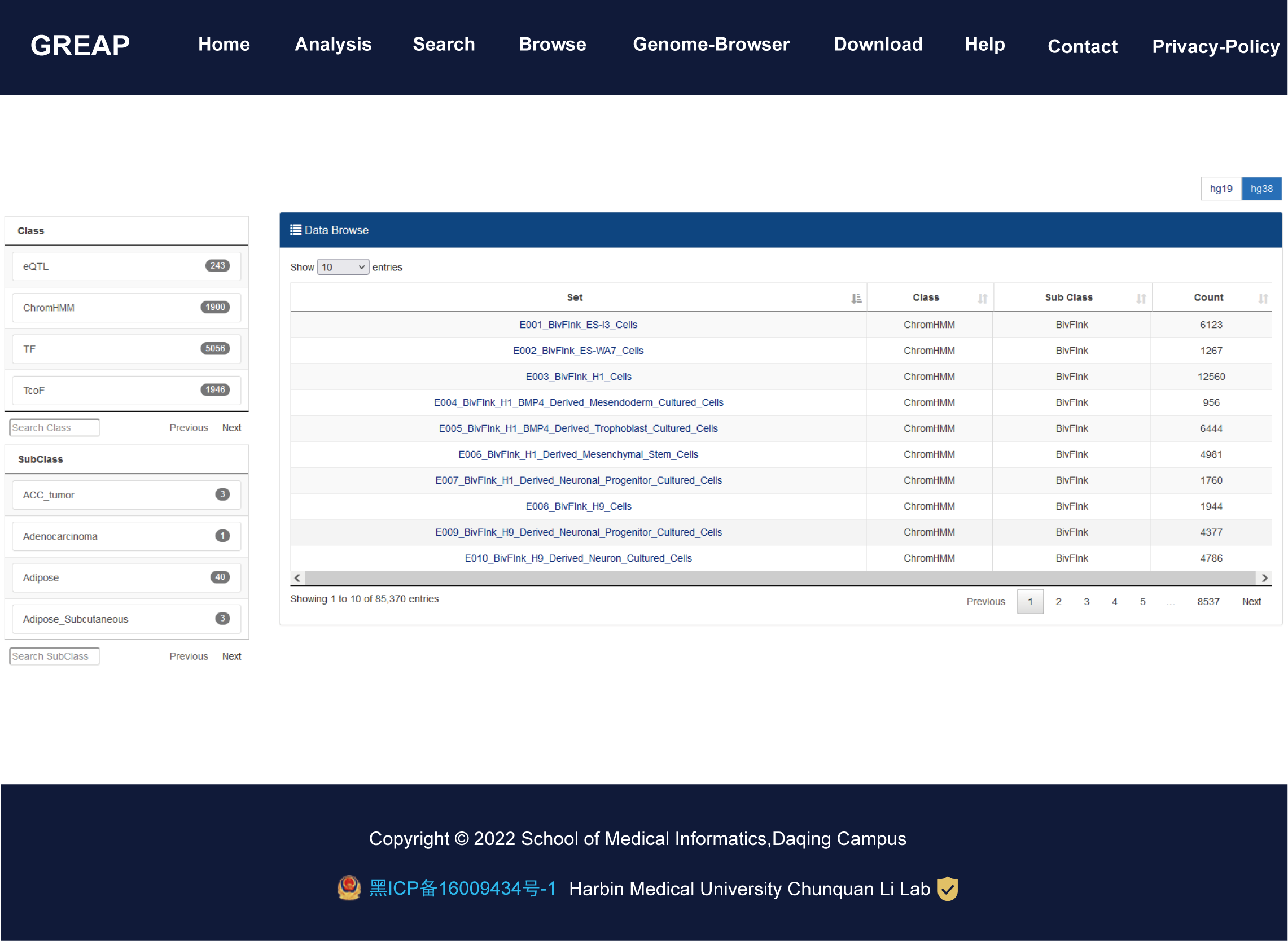
 Importantly, GREAP provides annotation and enrichment analysis functions of genomic region sets. In addition, GREAP provides a user-friendly interface to search, browse and visualize detailed information about these genomic region sets. In summary, GREAP is a powerful platform that provides a variety of types of genomic region sets for users and supports genomic region annotation and enrichment analysis functions.
Importantly, GREAP provides annotation and enrichment analysis functions of genomic region sets. In addition, GREAP provides a user-friendly interface to search, browse and visualize detailed information about these genomic region sets. In summary, GREAP is a powerful platform that provides a variety of types of genomic region sets for users and supports genomic region annotation and enrichment analysis functions.
| Data class | Set | Number of region |
| ChromHMM | 1,900 | 55,502,957 |
| TF | 5,056 | 54,910,288 |
| TcoF | 1,946 | 20,200,723 |
| Histone | 1,449 | 175,953,536 |
| ATAC | 2,986 | 51,964,468 |
| Enhancer | 543 | 6,621,919 |
| Super Enhancer | 641 | 396,631 |
| SNP | 11,652 | 34,986,456 |
| Methylation | 360 | 52,300,833 |
| LncRNA | 1,956 | 73,119 |
| mRNA | 5,6881 | 1,420,785 |
Principal Investigator:Chunquan Li, Ph.D.
Phone: 86-459-8153035
Fax: 86-459-8153035
Email: lcqbio@163.com
School of Medical Informatics,
Daqing Campus Harbin Medical University
39 Xinyang Road, Daqing 163319, China
SEdb
SEdb: the comprehensive human Super-Enhancer database
ENdb
ENdb: an experimentally supported enhancer database for human and mouse
SEanalysis
SEanalysis: a web tool for super-enhancer associated regulatory analysis
ATACdb
ATACdb: a comprehensive human chromatin accessibility database
KnockTF
KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription
factors
TRCirc
TRCirc: a resource for transcriptional regulation information of circRNAs
TRlnc
TRlnc: a comprehensive database of human transcriptional regulation of lncRNAs
VARAdb
VARAdb: a variation annotation database for human